
(© New Africa - stock.adobe.com)
Before the Fever Hits: A New Blood Test Could Warn Doctors Days in Advance
In A Nutshell
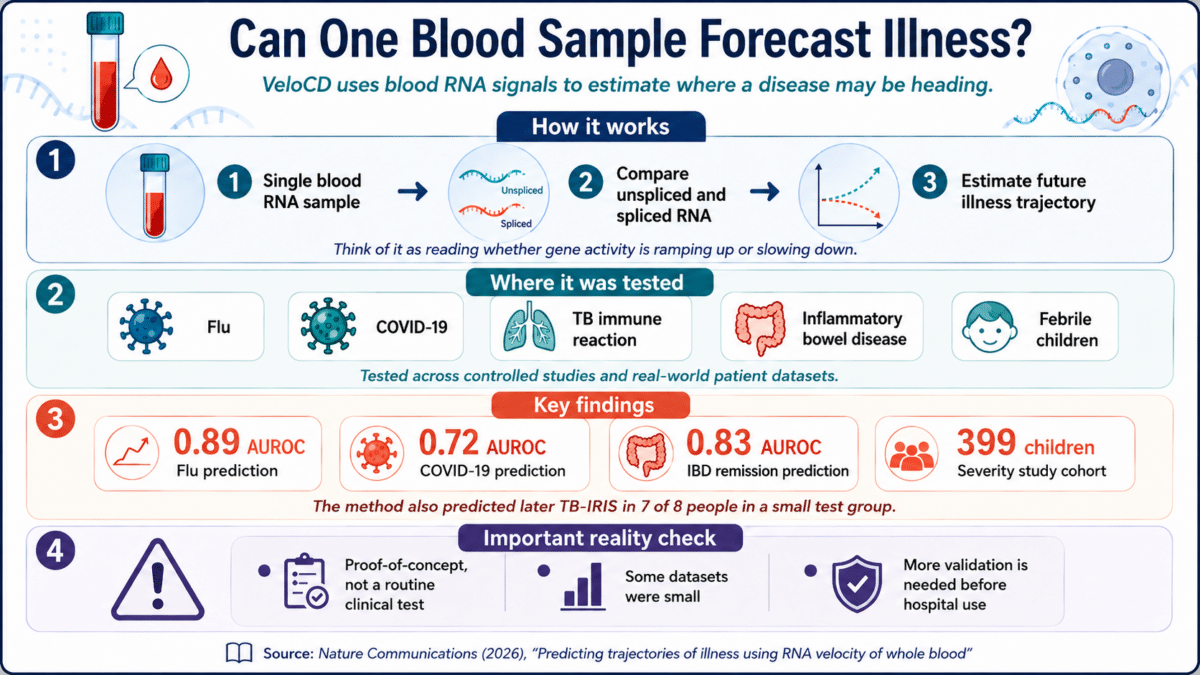
- Researchers developed VeloCD, a method that reads RNA activity in a single blood sample to estimate where a patient’s illness is headed before symptoms fully develop.
- In controlled studies, the tool predicted flu infection up to three days in advance and COVID-19 infection status up to 48 hours ahead.
- VeloCD was also tested across tuberculosis complications, inflammatory bowel disease treatment response, and severity of illness in 399 hospitalized children across Europe.
- The method is still proof-of-concept and not available for clinical use, but researchers say it could one day help doctors make faster, more informed decisions about who needs urgent care.
Scientists may have found a way to read the RNA signals in blood. Not just where an immune system stands right now, but where it is headed. Researchers developed a method called VeloCD that analyzes blood samples to forecast the future course of illnesses ranging from the flu and COVID-19 to life-threatening tuberculosis complications and severe infections in children.
RNA activity in blood cells can reveal whether key genes are ramping up or winding down, making it possible to estimate what a patient’s condition may look like hours or days from now. Researchers caution the method is still proof-of-concept, and significant further work is needed before it could reach everyday medical practice.
Blood contains not just finished RNA messages but also unfinished ones, raw and unprocessed, that hint at what cells are about to do next. By comparing these two layers, researchers can essentially read the direction an immune system is traveling, not just its current state. When a gene gets activated, raw unfinished material appears before the finished product is assembled and put to work. Most blood tests only measure the finished product. VeloCD measures both, and the ratio between them acts like a speedometer, showing whether production is accelerating or slowing for any given gene. VeloCD is not a commercially available test and was used only in controlled research settings.
Flu and COVID-19: Testing the Concept in Controlled Infections
Researchers started with carefully monitored experiments where healthy adult volunteers agreed to be exposed to a virus under medical supervision. In the influenza study, 17 of 23 subjects became infected; in the COVID-19 study, 17 of 26. Blood samples were collected at regular intervals before and after exposure.
Using blood samples taken just one day after exposure in the flu study, VeloCD produced an AUROC score of 0.89, a strong result on a standard scale where 1.0 is perfect. For COVID-19, the tool predicted infection status up to 48 hours in advance with an AUROC of 0.72. These numbers come from controlled experimental conditions and may not directly translate to real-world performance.
Three specific genes, including ones involved in how cells respond to viral threats and how fats move through the body, were enough to flag flu infection in blood drawn before symptoms appeared. Two of those same genes also ranked among the top predictors for COVID-19.
From Tuberculosis Complications to Bowel Disease
VeloCD’s reach extended beyond respiratory viruses, according to the Nature Communications study. In a clinical trial involving 95 patients co-infected with HIV and tuberculosis, blood samples were collected two weeks after patients started antiviral treatment. Some were at risk of developing a dangerous immune overreaction that can occur when an immune system begins recovering from HIV suppression and encounters tuberculosis bacteria. VeloCD correctly predicted which patients would develop this complication in 7 of 8 patients in the test group, with an AUROC of 0.72 in a broader formal test. The authors note the test group of 8 is small, making these results preliminary.
In a dataset of just 13 patients with inflammatory bowel disease, VeloCD used blood samples taken two weeks after the first infusion to predict which patients would be in remission by week 14. Despite the small sample, which the authors flag as a significant limitation, the tool achieved an AUROC of 0.83. Early signals from that cohort suggest the approach could potentially offer insight into whether a treatment is working before any clinical signs of improvement appear.
Blood RNA Signals and the Triage Problem in Sick Children
Perhaps the most immediately pressing application was in a multi-country study of 399 children admitted to hospitals across Europe with suspected infections. In real emergency settings, doctors face a difficult triage problem: most feverish children have mild illnesses they will shake off quickly, but some will deteriorate rapidly. Sending the wrong child home could be fatal.
Children were grouped by severity: mild (well enough to go home), moderate (admitted to a hospital ward), and severe (admitted to a pediatric intensive care unit). Using 59 genes identified through the analysis, VeloCD produced very different probability scores for children at opposite ends of the severity spectrum. Among mild illness cases in the test group, the median probability score for severe illness was extremely low. Among severe cases requiring immediate intensive-care transfer, the median score was 1.0, and most had scores above 0.75.
Children with moderate illness were harder to read, and scores varied more widely, as expected. Among those children, the tool’s ability to predict later intensive-care transfer was modest, with an AUROC of 0.72, and was not clearly better than CRP, a standard inflammation blood marker, in the available comparison. Still, children with higher probability scores were more likely to have elevated inflammation markers, more likely to require surgical procedures, and more likely to stay in the hospital longer.
A blood-based RNA test that estimates where an illness may be heading from a single sample would mark a real shift in how medicine approaches some of its hardest problems. VeloCD has real promise and real limitations. But the core idea, that blood is already broadcasting early signals about what is coming next, is becoming harder to dismiss.
Disclaimer: This article is based on a peer-reviewed study. The research described is proof-of-concept and has not been validated for clinical use. VeloCD is not an approved diagnostic tool and is not available to patients or physicians. Findings from small study cohorts should be interpreted with caution.
Paper Notes
Limitations
The authors describe their findings as proof-of-concept and acknowledge several important limitations. Several datasets were small and unbalanced, particularly the inflammatory bowel disease cohort of 13 patients and the tuberculosis immune reconstitution inflammatory syndrome test group of only 8 subjects, which limits the strength of conclusions that can be drawn. Performance varied depending on technical settings, meaning results were not uniform across all analysis configurations. In the febrile children study, most moderate-illness children received treatments determined by their attending physicians, which likely reduced the chance of deterioration and limited the tool’s ability to make categorical predictions in that group. The authors also noted that further work may be needed to optimize reference datasets for different disease contexts, and that VeloCD works best when genes are carefully selected to distinguish between disease states.
Funding and Disclosures
Funding was provided by multiple sources including the European Union’s Horizon 2020 Research and Innovation programme (grant agreement numbers 668303 and 848196), the UK Medical Research Council, the UK Engineering and Physical Sciences Research Council, the NIHR Imperial Biomedical Research Centre, the Wellcome Trust, the Francis Crick Institute, the US Defense Advanced Research Projects Agency, the South African Research Chairs Initiative, and Taiwan’s National Science and Technology Council, among others. An international patent application for a method to predict disease prognosis based on RNA velocity has been filed by Imperial College Innovations Limited, with authors A.J. Cunnington, C. Dunican, M. Barahona, and M. Kaforou listed as inventors. Other authors declared no relevant competing interests.
Publication Details
Paper title: Predicting trajectories of illness using RNA velocity of whole blood. Published in Nature Communications, volume 17, article number 3652, 2026. DOI: https://doi.org/10.1038/s41467-026-71685-5. Received January 8, 2025; accepted March 26, 2026; published online May 6, 2026. Corresponding authors: Claire Dunican and Aubrey J. Cunnington, Department of Infectious Disease, Imperial College London. A full author list and institutional affiliations appear at the end of the published article.







