
(Photo by Kyrylo Ryzhov on Shutterstock)
In A Nutshell
- Researchers found that some bacteria common in the mouth also appear in the gut of stomach cancer patients.
- Genetic comparisons showed many of these bacteria closely match within the same person, suggesting oral-to-gut transmission.
- Saliva and stool samples could help detect patterns linked to stomach cancer, but tests are not ready for clinical use.
- The study cannot show whether these bacteria cause cancer or simply thrive once it develops.
Bacteria that live in the mouth don’t always stay there. Some slip down into the gut and make themselves at home. A large new genetic study has found that specific mouth bacteria migrate to the gut and thrive in people with stomach cancer, and that a simple saliva or stool sample might help flag patterns linked to the disease without an invasive scope down the throat.
The research analyzed genetic material from stool and saliva samples of 317 people across two separate patient groups, with additional testing in a third group from Harbin, China.
A clear pattern emerged: a family of acid-producing bacteria that normally live in the mouth were found growing in abundance in the guts of stomach cancer patients, a pattern distinct from what researchers saw in people with chronic gastritis, a less severe stomach condition. Strain-level genetic fingerprinting revealed near-identical bacterial strains in many mouth-gut comparisons within the same person, pointing to direct transmission along what the researchers describe as an “oral-gastric-gut” pathway of disease.
Researchers also built machine-learning models that could predict whether someone had stomach cancer based solely on their bacterial signatures. The saliva-based model achieved an AUROC, a standard measure of how well a diagnostic test distinguishes between people with and without a disease (where 1.0 is perfect and 0.5 is no better than chance), of 0.87, while the stool-based model reached 0.85. Those scores suggest potential for future noninvasive detection tools for a major global cancer whose burden falls heavily on East Asia.
How the Study Worked
Scientists from BGI Genomics and Shanghai Jiao Tong University’s Renji Hospital collected stool and saliva samples from patients diagnosed with either stomach cancer or chronic gastritis. A technique that shreds all the genetic material in a sample into tiny fragments and reads them with powerful sequencing machines produced a detailed, species-level snapshot of every microbe present.
Cohort one included 106 people with chronic gastritis and 106 with stomach cancer, all providing stool samples. Cohort two had 52 chronic gastritis patients and 53 stomach cancer patients, providing both stool and saliva samples. A third, independent group from Harbin provided tongue swab samples for further testing. Eighty-seven patients had matched stool and saliva samples, allowing researchers to directly compare the bacteria living in each person’s mouth with those in their gut.
After sequencing, the team identified 28 bacterial species in stool that reliably differed between cancer and non-cancer patients across both main groups. Of these, 23 species were more abundant in stomach cancer patients. An overwhelming majority, 20 out of 28, were species found in both the mouth and gut. Most belonged to types of bacteria that produce lactic acid as a byproduct, organisms that are typically harmless or even beneficial in the mouth but that appear problematic when they accumulate in large numbers in the gut. Eight of these species belonged to Streptococcus, a group commonly associated with the mouth.
Tracing Mouth Bacteria to the Gut in Stomach Cancer Patients
One of the study’s most important contributions was showing, at the genetic strain level, that bacteria in the mouth and gut weren’t just similar; in many within-person comparisons, they were near-identical. Researchers compared DNA sequences of bacteria found in different body sites within the same person. For eight Streptococcus species examined, between 42% and 83% of matched mouth-gut comparisons within the same person showed near-perfect genetic matches. When the team compared saliva bacteria between different people, fewer than 3% of comparisons reached that same threshold. In other words, the findings strongly suggest the gut bacteria originated from each patient’s own mouth rather than from some shared environmental source. Age was not a major factor in these patterns, based on the team’s analysis.
Beyond identifying which bacteria were present, the team investigated what those bacteria were doing. Analysis of the genetic data revealed that metabolic pathways involved in lactic acid production were far more active in the guts of stomach cancer patients. All seven enzymes involved in one fermentation pathway were elevated in cancer patients from the first group, including enzymes directly responsible for producing lactic acid. Heightened lactic acid production matters because it can create an acidic environment around tumors that prior research has linked to cancer spread, the growth of new blood vessels feeding tumors, and a weakened immune response against cancer cells.
Researchers propose what they call a “transmission to colonization” model, a hypothesis, not yet proven, about how this process might work. The idea is that stomach cancer creates a welcoming environment (lower stomach acid, altered immune defenses, and new nutrient sources from the tumor itself) that allows mouth bacteria to survive a journey they normally wouldn’t and then set up shop in the gut. Rather than acting as random hitchhikers, these bacteria may actively contribute to cancer progression by producing lactic acid and forming cooperative communities that help each other survive.
Cancer-associated bacteria weren’t operating alone, either. The lactic acid bacteria enriched in cancer patients formed tightly connected cooperative networks in both the mouth and gut. Several Lactobacillus and Streptococcus species clustered together with other acid-producing bacteria, suggesting these organisms may work as a team to colonize new territory and resist the body’s defenses. Among the shared species enriched in stomach cancer, nearly 75% of all possible bacterial pairings showed significant relationships in one dataset, compared to roughly 20% for other shared species.
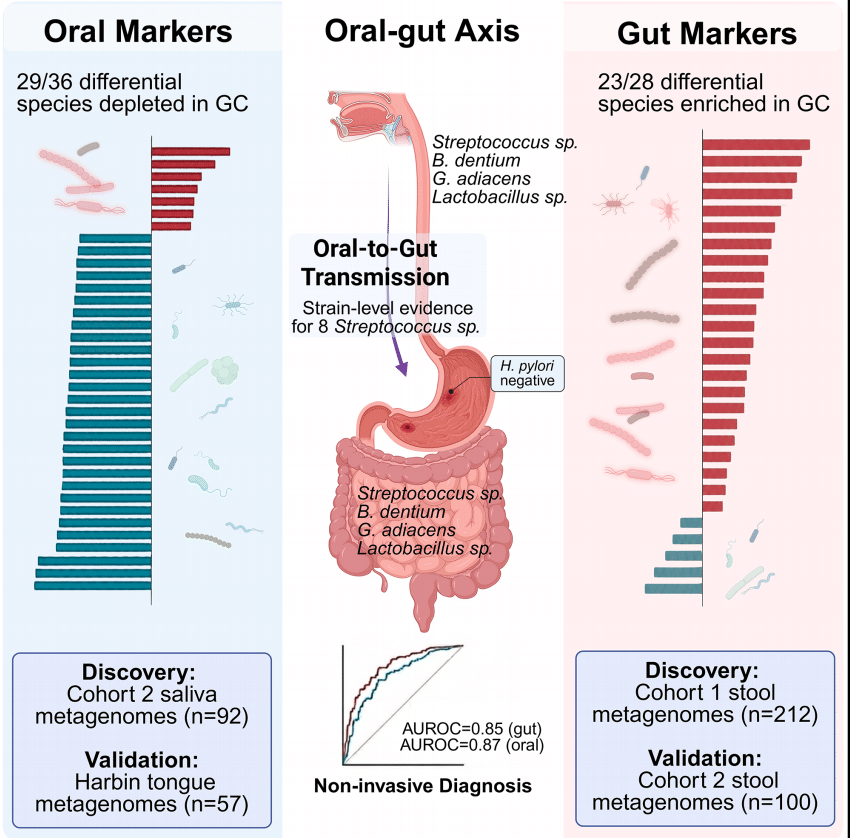
• Streptococcus species transmission from mouth to gut confirmed at the strain level
• Microbiome classifiers enable accurate noninvasive GC detection
Credit: BGI Genomics
Could a Saliva Test Screen for Stomach Cancer?
Predictive models the team built are worth a closer look. For stool samples, researchers identified 17 bacterial markers, 10 of which were mouth-to-gut transmitters, that a machine-learning algorithm used to predict cancer status with an AUROC of 0.85. For saliva, 20 markers fed a model that achieved an AUROC of 0.87 when tested against the independent Harbin tongue samples. Notably, the saliva model outperformed the stool model, which the researchers suggest could mean changes in the mouth appear earlier or are more pronounced than gut changes in stomach cancer.
A saliva test that could one day help screen for stomach cancer is especially appealing in parts of the world where the disease is most common and endoscopy, the standard clinical method, remains the main diagnostic tool.
Caveats remain. Because the study captured a single snapshot in time rather than tracking patients over years, what scientists call a cross-sectional design, it cannot prove that these bacterial changes drive cancer rather than result from it. Untangling cause from effect will require long-term follow-up studies and animal experiments, both of which the authors call for. Most patients in the study were also negative for H. pylori, the bacterium long established as the primary risk factor for stomach cancer, meaning the findings may not apply equally to infected populations.
What the study, published in Cell Reports Medicine, does establish, across multiple patient groups and body sites, is that the mouth and gut are not separate microbial kingdoms. Traffic between them matters in stomach cancer. Whether these traveling bacteria turn out to be drivers or passengers, they leave a signature distinctive enough to change how this deadly disease might one day be detected.
Paper Notes
Limitations
Several caveats were acknowledged by the authors. Most analyzed samples came from patients who were negative for H. pylori, so findings may not generalize to H. pylori-positive stomach cancer patients. Researchers did not assess participants’ oral health status, such as the presence of cavities, which could affect oral-gut transmission patterns. Their sequencing approach prioritized bacterial communities, meaning fungal and viral contributions were under-represented. Saliva sample size was relatively modest at 92 samples, and saliva may not fully capture the diversity of different oral areas such as the tongue, inner cheek, or dental plaque. A cross-sectional design, measuring everything at one point in time, prevents researchers from determining whether the bacterial enrichment causes or results from stomach cancer progression. Stool microbiome composition is also influenced by multiple organs beyond the stomach and gut, including the liver and pancreas, adding potential confounding factors.
Funding and Disclosures
Support came from the State Key R&D Program (2020YFA0509200), Noncommunicable Chronic Diseases–National Science and Technology Major Project (2025ZD0551704), National Natural Science Foundation of China (82530086, 82330086, 82203224, and 82570645), Shanghai Jiao Tong University 2030 Initiative (WH510272101), Heilongjiang Provincial Key Research and Development Program (no. 2023ZX07D05), Shanghai Rising-Star Program (24QA2705000), Shanghai Eastern Talent Plan Youth Project (QNWS2025114), Chenguang Program of Shanghai Education Development Foundation and Shanghai Municipal Education Commission (22CGA17), and Shanghai Youth Medical Talents – Specialist Program (2023-62). Authors declared no competing interests. During manuscript preparation, the authors used ChatGPT, Grok, Gemini, and DeepSeek to polish English writing; all authors reviewed and take full responsibility for the content.
Publication Details
Title: Distinct signatures in the human gut and oral microbiomes of gastric cancer | Authors: Youwen Qin, Ya-Xuan Zhang, Li-Ping Liu, Yuan-Hong Xie, Xin-Yue Ma, Yu Hao, Li-Cong Zhao, Jiao-Jiao Dong, Ying He, Ke Sun, Huanzi Zhong, Shida Zhu, Ming Liu, Jing-Yuan Fang, and Cheng-Bei Zhou | Affiliations include: Shenzhen Hospital of Southern Medical University; BGI Genomics (Shenzhen and Harbin); BGI Research, Shenzhen; Division of Gastroenterology and Hepatology, Renji Hospital, School of Medicine, Shanghai Jiao Tong University; China-Japan Friendship Hospital, Beijing; Fourth Affiliated Hospital of Harbin Medical University; Harbin Fourth Hospital; Harbin Sixth Hospital | Journal: Cell Reports Medicine, Volume 7, 102761, May 19, 2026 | DOI: 10.1016/j.xcrm.2026.102761 | Data Availability: Raw sequencing data deposited in CNGB Sequence Archive (CNSA) under accession number CNP0004342. Code available at GitHub: https://github.com/Owen-haha/GC_OralGut_MWAS







