
UCT, Glycobiomedical Laboratory. Credit: Scientific Computing Research Unit (SCRU)
In A Nutshell
- Cancer cells produce distinctive sugar markers on a protein called MUC1 by relocating key enzymes to the wrong compartment inside the cell, giving them unchecked time to work without competition.
- The findings could help guide the design of more precisely targeted cancer vaccines and therapies, though further validation in living systems will be needed.
- A University of Cape Town team built a laboratory model that recreates both healthy and tumor cell sugar-building conditions, mapping exactly which enzymes act on which sites and in what order.
Experimental cancer vaccines targeting a molecule called the sTn antigen have been in development for years. Researchers have largely been shooting at a moving target, however, unsure of precisely where on a key cancer protein that antigen actually appears. A new study identifies its most likely molecular address, a finding that could help guide how the next generation of cancer vaccines and targeted therapies gets designed.
At the center of this work is a protein called MUC1, short for mucin 1. Every cell in the body wears a coat of sugar molecules on its surface, and MUC1 is one of the proteins that carries that coat. In healthy tissue, MUC1’s sugars are arranged in a carefully ordered pattern, assembled by specialized enzymes working in sequence. In cancer, that pattern gets scrambled, and the result is a distinctive set of abnormal sugar structures that doctors already use as tumor markers. One of the most clinically tracked is the sTn antigen, which shows up across a wide range of cancer types and has been a target for vaccine developers for decades. Until now, the field lacked a precise map of where on MUC1 that antigen is most likely to land.
Researchers at the University of Cape Town have built that map, and in doing so, uncovered the step-by-step cellular mechanism behind it.
How Cancer Changes MUC1’s Sugar Coat
A team led by Kevin J. Naidoo developed what they call a “one-pot” laboratory approach. It’s essentially a simplified system using a single reaction vessel to recreate the sugar-building assembly line normally spread across multiple compartments inside a living cell. Their system let them track which enzymes act on which specific spots along the MUC1 protein, how competition between enzymes changes the final product, and how moving one group of enzymes to the wrong cellular address (something that happens in real tumors) produces the cancer-associated sugar signatures doctors look for in patient tissue.
Cells build sugar coatings on proteins through a relay system spanning two major internal compartments. One is a sprawling network of membranes called the endoplasmic reticulum, essentially the cell’s protein factory, where proteins are first assembled. The other is a series of stacked compartments called the Golgi apparatus, where proteins receive their final sugar decorations before reaching the cell surface.
In healthy cells, the first sugar gets attached to MUC1 by a family of enzymes called GALNTs, which normally operate in the Golgi. Once that initial sugar is placed, other enzymes add more on top, building out chains in a strict sequence. In cancer cells, the GALNT enzymes get relocated backward into the earlier compartment away from where they belong and away from the competing enzymes that would normally keep them in check. Previous research on breast cancer tissue found that a particular sugar marker associated with this relocation was 4.5-fold more abundant in tumor samples than in normal tissue, and that 70 percent of high-expression samples showed this backward shift of GALNTs.
Building a Tumor’s Sugar Pattern in a Test Tube
To model this, the researchers attached a MUC1 protein sequence to a carrier molecule that could be produced in bacteria, purified, and run through controlled enzyme reactions. They confirmed the carrier didn’t interfere with any enzyme’s natural behavior. Reaction rates were identical with or without it attached.
Running experiments with four GALNT enzymes, both individually and in combination, the team mapped all five sugar-attachment sites on MUC1. Two of them, called GALNT1 and GALNT2, could independently place the first sugar at certain sites by latching directly onto the bare protein. Two other sites were handled exclusively by a third enzyme, GALNT4, which could only act after the first two had already placed sugars nearby. It uses a built-in sugar-sensing region to find its targets rather than working from scratch. A fourth enzyme tested showed no activity on any sequence at all.
When the researchers simulated healthy cellular conditions, running GALNT enzymes alongside C1GALT1, a competing enzyme that normally operates alongside them in the Golgi, sugar-site occupancy was limited. That competing enzyme quickly converted freshly placed sugars into a form the GALNTs’ sensors could no longer read, effectively braking the process. Only three of the five sites got decorated, matching what scientists observe in healthy tissue.
Simulating cancer’s enzyme relocation told a different story. When the GALNT enzymes were allowed to work first, in isolation and without competition, before the competing enzyme was introduced afterward, all five sites became fully occupied in the experimental system.
Introducing ST6GALNAC1, the enzyme responsible for producing the sTn cancer marker: in place of the competing enzyme at that stage produced the sTn antigen at every available site, the hallmark pattern of tumor-associated MUC1.
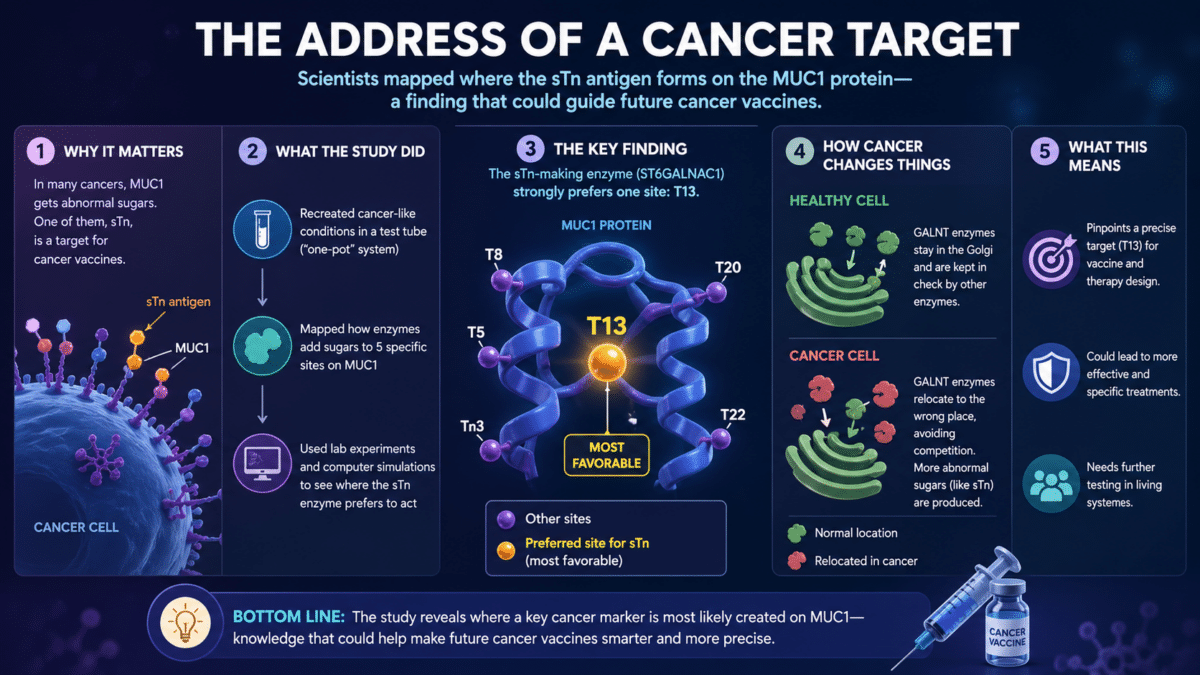
A Preferred Vaccine Target Emerges: Why T13 Matters
That last finding set up the study’s most clinically relevant result. With ST6GALNAC1 identified in this model as the primary enzyme producing sTn on MUC1, the researchers used a combination of lab experiments and atomic-level computer simulations to determine which of the five sites it prefers.
The simulations, published in Nature Communications, calculated how much chemical energy ST6GALNAC1 must expend to complete its reaction at each candidate site: essentially, how high a hurdle it has to clear. T13, a specific amino acid building block sitting within a region of MUC1 called PDTR, presented the lowest hurdles at both steps of the reaction, making it the most favorable target. A site called T8 presented the highest hurdles, making it the least favorable. The enzyme also showed essentially no activity on a different category of amino acid building blocks at those positions, sticking almost exclusively to the type found at T13.
Lab analysis of the actual reaction products, using a technique that identifies molecular structures by their mass, supported the computational picture: the sTn-producing enzyme was active at T13, T8, and T20, with T13 emerging as the preferred site. A related enzyme, ST6GALNAC2, has sometimes been linked to sTn production in cancer. It showed no significant activity on MUC1 at all, identifying ST6GALNAC1 as the primary driver of sTn formation on this protein, at least within the model system studied.
For vaccine developers, the precision matters. The sTn antigen is already a target in several experimental immunotherapy programs. Knowing that it appears predominantly at the T13 position on MUC1 rather than being distributed unpredictably across the protein could help researchers design more precisely targeted treatments, even if further validation in living systems will be needed.
A Mechanism, Not Just a Map
The T13 finding doesn’t stand alone. It follows directly from the broader mechanism the study laid out: in healthy cells, enzyme competition in the Golgi limits sugar-site coverage and keeps the sites that require a sugar-sensing relay, including T13, largely unoccupied. In tumor cells, the GALNT enzymes relocate to the protein factory, work without interference, fill all five sites including T13, and leave that site primed for the sTn-producing enzyme to install its cancer marker.
That chain of events, from enzyme misplacement to cancer marker at a preferred molecular location, reconstructed here in a controlled laboratory model, had not previously been mapped in this level of mechanistic detail. For researchers building diagnostics or therapies around MUC1’s tumor-associated sugar structures, the question of where those structures appear and how they get there now has a concrete, mechanistic answer.
Disclaimer: This article describes early-stage laboratory research, not an approved cancer vaccine or treatment. The work still requires further validation in living systems and clinical studies. Nothing in this article should be taken as medical advice, a diagnosis, or a recommendation for treatment. Anyone with questions about cancer prevention, diagnosis, vaccines, or treatment options should consult a qualified healthcare professional.
Paper Notes
Limitations
The study acknowledges several limitations. Measurements reported are described as “apparent values” because the protein sequences contain multiple reactive sites, meaning the measurements reflect the combined contribution of all sites rather than individual site behavior. The researchers also note that enzyme performance under real-world cellular conditions is influenced by factors including enzyme attachment to membranes, responses to more involved protein structures in living cells, and shifting concentrations of enzymes and their targets — none of which are fully captured in a test-tube system. The computer simulations are described as providing comparative, qualitative insight rather than absolute measurements. The authors note that the single-repeat model cannot account for the influence of neighboring repeats on sugar attachment in the full-length MUC1 protein found in nature. Separating certain effects from one another would require additional investigation involving genetic modification of the enzymes and more advanced analysis methods, which the authors state was outside the scope of this work. LC-MS samples were each analyzed once, with no technical or biological replicates, as samples were obtained from a single in vitro synthesis batch.
Funding and Disclosures
This work was supported in part by the National Research Foundation (NRF) CPPR 466624 grant and the South African Medical Research Council Self-Initiated Grant (SAMRC SIG) 416090. Graduate fellowship funding was provided by the Scientific Computing Research Unit (SCRU) at the University of Cape Town, and computational resources were provided by the Center for High Performance Computing (CHPC). Corresponding author Kevin J. Naidoo discloses that structural data identified through QM/MM simulations in this work have potential commercial applications, including drug design and development, and that he intends to pursue intellectual property protection on that data. Naidoo also holds shares in Struenodo, which provided experimental materials free of charge. All materials used are commercially available. The remaining authors declare no competing interests.
Publication Details
Title: An in vitro approach for simulating divergent Golgi O-glycosylation of tumor-associated MUC1 from normal MUC1 | Authors: Abdullateef Nashed, Kyllen Dilsook, Tharindu Senapathi, and Kevin J. Naidoo | Affiliations: Scientific Computing Research Unit, University of Cape Town, Rondebosch, South Africa; Department of Chemistry, University of Cape Town, Rondebosch, South Africa; Department of Chemistry, Faculty of Applied Sciences, University of Sri Jayewardenepura, Nugegoda, Sri Lanka | Journal: Nature Communications (2026), Volume 17, Article 3619 |DOI: 10.1038/s41467-026-72151-y | Received: 23 January 2025 | Accepted: 6 April 2026 | Published online: 22 April 2026 | Corresponding author: Kevin J. Naidoo ([email protected])







