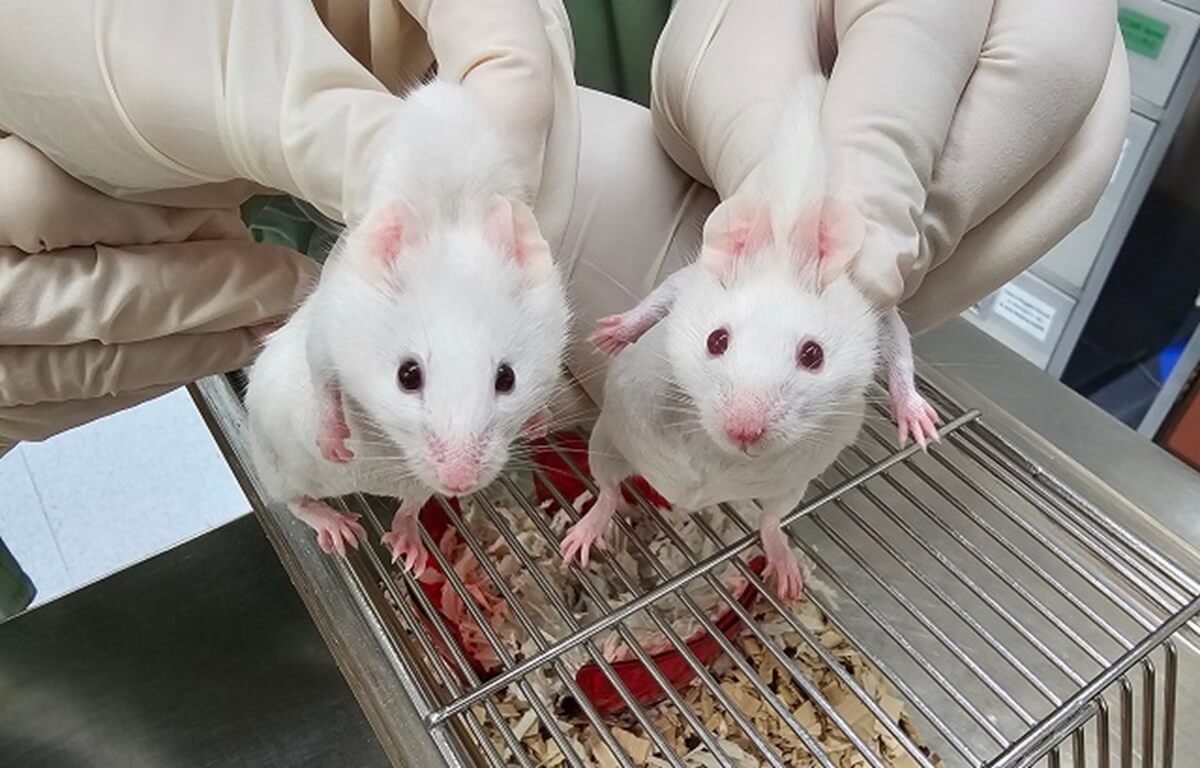
The mouse on the left is a chimeric with dark eyes and patches of black fur, a result of stem cells derived from a choanoflagellate Sox gene. The wildtype mouse on the right has red eyes and all white fur. The color difference is due to genetic markers used to distinguish the stem cells, not a direct effect of the gene itself. Credit: Gao Ya and Alvin Kin Shing Lee, with thanks to the Centre for Comparative Medicine Research (CCMR) for their support. (Credit: Queen Mary University of London)
LONDON — Could genes from the dawn of life hold the key to a stem cell revolution? A groundbreaking experiment could make it reality. Researchers have successfully created a living mouse using genetic tools from a microscopic organism that lived nearly a billion years ago. This scientific feat not only challenges our understanding of cellular evolution but also opens new doors for regenerative medicine.
Scientists from Queen Mary University of London and The University of Hong Kong used genes from choanoflagellates — tiny, single-celled organisms that are the closest living relatives to animals — to generate stem cells capable of developing into a complete mouse. By introducing ancient genes into mouse cells, the researchers demonstrated an extraordinary continuity of genetic function across hundreds of millions of years of evolution.
“By successfully creating a mouse using molecular tools derived from our single-celled relatives, we’re witnessing an extraordinary continuity of function across nearly a billion years of evolution,” says Dr. Alex de Mendoza from Queen Mary University of London, the lead researcher of the study published in Nature Communications, in a media release.
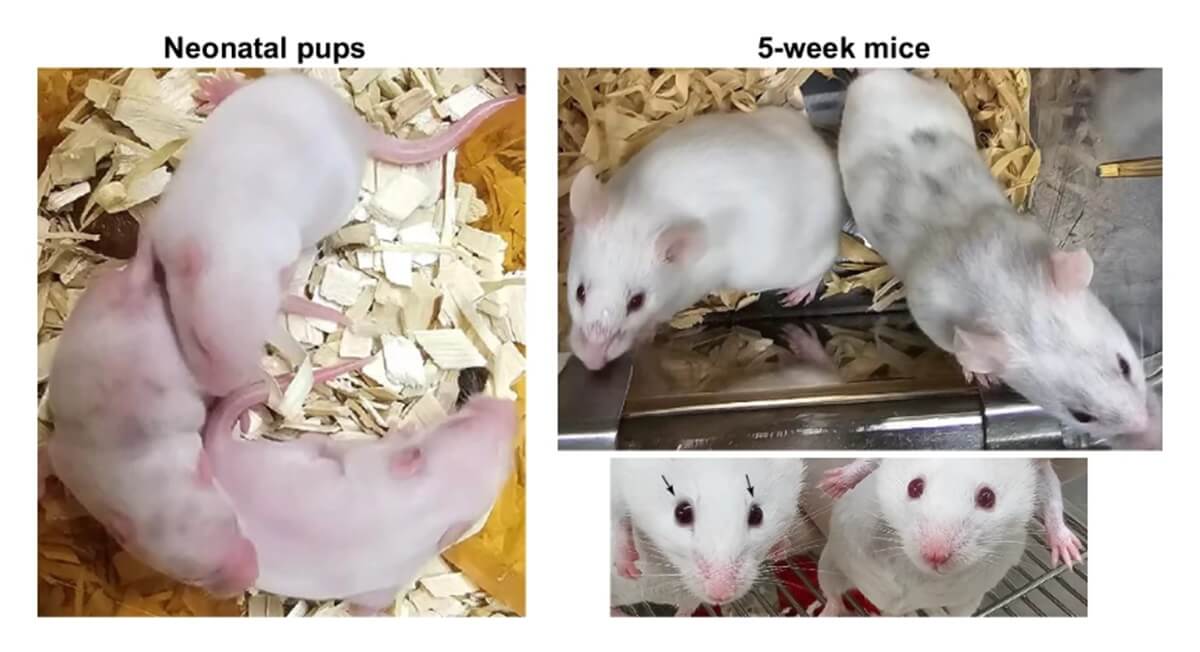

The experiment involved replacing a mouse’s native Sox2 gene with a similar gene from a choanoflagellate. When these genetically modified cells were injected into a developing mouse embryo, they successfully integrated, producing a chimeric mouse with distinctive traits like black fur patches and dark eyes.
What makes this discovery particularly fascinating is its implications for understanding how complex life might have emerged. The choanoflagellate genes, which were likely originally used to control basic cellular processes, were repurposed by multicellular organisms to drive stem cell formation and development.
Beyond its evolutionary significance, the research could revolutionize regenerative medicine. By exploring these ancient genetic tools, scientists might develop more effective stem cell therapies and advanced techniques for treating diseases or repairing damaged tissues.
“Studying the ancient roots of these genetic tools lets us innovate with a clearer view of how pluripotency mechanisms can be tweaked or optimized,” Dr. Ralf Jauch from The University of Hong Kong notes.
This breakthrough reminds us that the building blocks of life are more adaptable and interconnected than we ever imagined, with tiny, ancient organisms holding profound secrets about our biological origins.
Paper Summary
Methodology
This study looked into the origins of two proteins, Sox and POU, that are vital for stem cells in animals. Researchers explored how these proteins work by studying their structure and functions in both animals and their unicellular relatives, such as choanoflagellates. Using experiments, they tested whether these proteins from single-celled organisms could perform similar tasks as their counterparts in animals. For example, they tried to reprogram mouse cells into stem cells using these proteins. They also recreated ancient versions of these proteins to see how their functions evolved over time.
Key Results
The researchers discovered that some unicellular organisms, related to animals, already have proteins like Sox and POU. These proteins can bind to DNA and, in some cases, even mimic the functions of animal stem cell proteins. For instance, certain choanoflagellate Sox proteins were able to replace their animal counterparts and successfully reprogram mouse cells into stem cells. However, the POU proteins from these unicellular organisms lacked this ability, showing a difference in their evolutionary roles.
Study Limitations
- Unicellular Sampling Gaps: The study relied on limited genomic data from unicellular organisms, which might not provide a complete picture.
- Functional Validation: The experiments showed some functional similarities but couldn’t fully replicate natural conditions of stem cell creation in animals.
- Context-Specific Evolution: While some functions were conserved, others might have evolved uniquely in animals, limiting the study’s ability to generalize findings across all lineages.
Discussion & Takeaways
The study highlights that critical proteins for animal stem cells, such as Sox and POU, existed even before animals evolved. These proteins played roles in simpler organisms, suggesting that their DNA-binding and regulatory abilities were repurposed for multicellular life. This evolutionary repurposing might explain how complex traits, like stem cells, emerged in animals. The findings also underline the deep evolutionary roots of some of the most essential biological mechanisms.
Funding & Disclosures
The research was funded by institutions like the University of Hong Kong and the Max Planck Institute for Terrestrial Microbiology. The authors declare no conflicts of interest. Ethical guidelines for animal use were followed as per the approvals of respective institutional boards.








You should contact these people and tell them to test the mice’s intelligence.