
DNA sequence (Credit: Gio.tto on Shutterstock)
We’re all mostly human. 8% of your genome comes from viruses.
In A Nutshell
- Scientists captured the first detailed images of HERV-K, a viral protein found in cancer cells, lupus, and ALS patients
- HERV-K comes from a virus that infected human ancestors millions of years ago; these ancient viral sequences make up 8% of human DNA
- The protein looks completely different from HIV and has unique structural features that could be important for developing treatments
- Antibodies created by researchers light up diseased cells from lupus and arthritis patients but ignore healthy cells
- Early experiments show promise for targeting HERV-K with antibodies and engineered immune cells to treat cancer
A protein showing up in breast cancer tumors, lupus patients, and people with ALS has been traced to an unexpected source: a virus that infected human ancestors millions of years ago. Scientists have now captured the first detailed images of this disease-linked protein, revealing what this genetic remnant actually looks like. These ancient viral sequences make up about 8% of the human genome.
The protein comes from HERV-K (Human Endogenous Retrovirus K), a retrovirus that infected primates millions of years ago and was permanently copied into their DNA. That viral DNA has been passed down through generations ever since. In healthy people, cells keep this ancient code locked away and silent. But something goes wrong in certain diseases. Cancer cells in breast, ovarian, prostate, and other tumors start making the HERV-K protein. Immune cells from patients with rheumatoid arthritis and lupus show elevated levels. In people with ALS, the protein floats in spinal fluid where it appears to damage neurons.
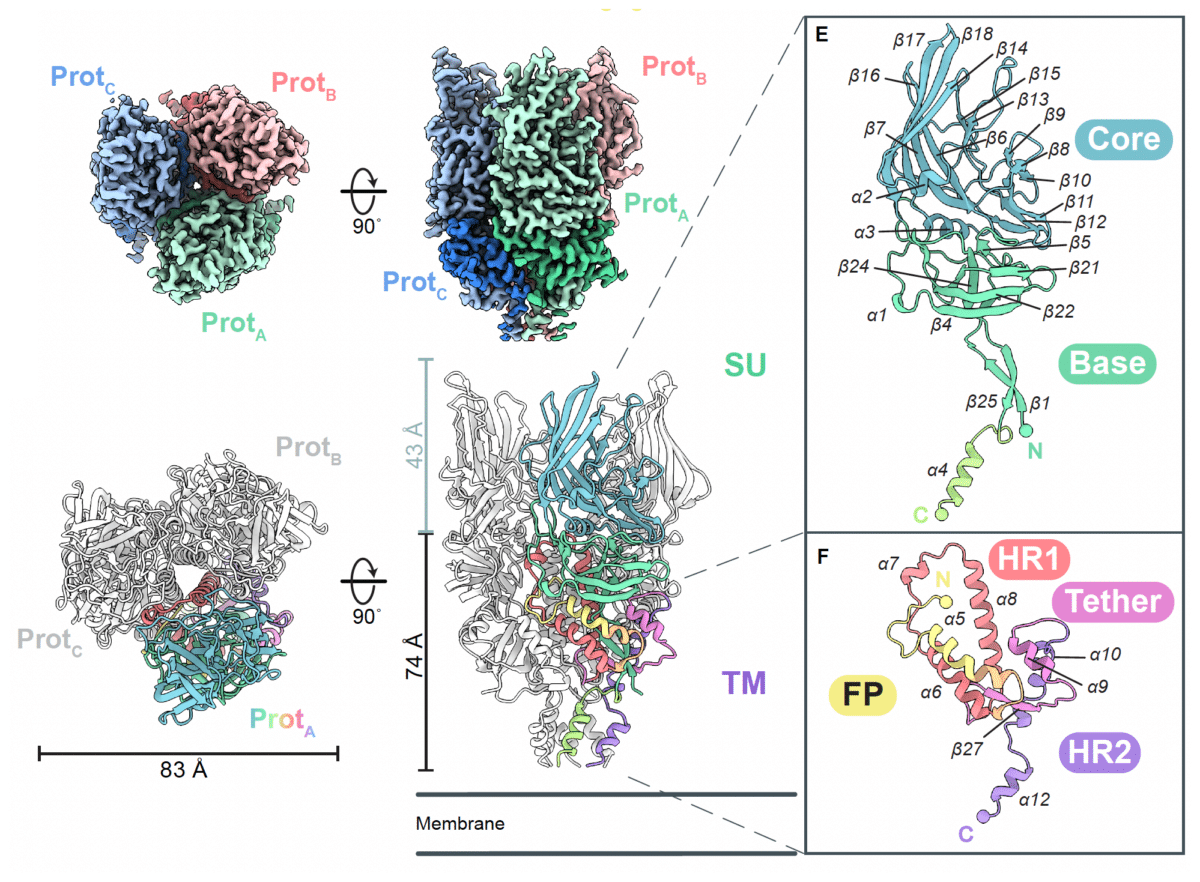
For decades, researchers have known this fossilized virus shows up in diseased tissue, but they’ve never seen what the protein looks like or understood how it works. The new study from the La Jolla Institute for Immunology, published in Science Advances, finally provides that picture. The images reveal the protein in both its ready-to-spring form and its activated state.
Catching a Shape-Shifter
The HERV-K protein naturally exists in an unstable state, like a mousetrap set and ready to snap. To take pictures, researchers needed to freeze it in that cocked position, but it kept collapsing into its sprung form before they could capture an image.
Lead researchers Jeremy Shek and Chen Sun engineered hundreds of modified versions, testing different ways to hold the protein in place. The winning approach worked like putting a safety pin in a grenade—they added molecular locks that kept the structure stable long enough to photograph. These stabilized versions could survive freezing and held together even under harsh conditions that would normally trigger the shape change.
HERV-K Shaped Differently from HIV
The images revealed something unexpected. HERV-K looks completely different from HIV and other well-studied viruses. It’s tall and narrow, shaped like an inverted tripod with three prongs. Each prong has two parts—a top section that latches onto cells and a lower section that handles the membrane fusion.
When the protein activates, this arrangement explodes into action. Part of it shoots out like a harpoon toward target cells, the whole structure extends like a telescope, then snaps back into a hairpin shape that forces two cell membranes together.
The activated form revealed something researchers hadn’t seen before: an extra structural piece sitting between two standard regions. This feature appears specific to HERV-K’s family of viruses and doesn’t exist in HIV or related viruses. That difference might be important for developing targeted treatments.
Antibodies Light Up Diseased Cells
To study HERV-K in real patients, researchers needed tools that could detect the protein and tell the difference between its various forms. The team created a set of antibodies—immune molecules that latch onto specific targets—by immunizing mice and screening which ones stuck to HERV-K.
Two antibodies proved especially useful for research. One recognizes only the ready-to-spring form of the protein, while another only grabs onto the activated version. These became crucial tools for confirming they had captured the right structures in the lab.
The real test came with patient samples. When researchers exposed immune cells from people with lupus and rheumatoid arthritis to five of the antibodies, the diseased cells lit up with fluorescent markers. The staining appeared primarily inside the cells. Notably, the antibody that only recognizes the activated form did not react—suggesting that patient cells contain the ready-to-spring version or other forms of the protein, not the activated state. Immune cells from healthy people showed no reaction at all.
HERV-K Opens Doors to New Treatments
The structures and antibodies create opportunities for targeting HERV-K in disease. Since the protein appears on cancer cell surfaces, it could serve as a target for immune-based therapies.
Previous studies have shown promise with this approach. One antibody inhibited breast cancer cell growth in laboratory tests and in tumor-bearing mice. Other researchers engineered immune cells to recognize and attack anything displaying HERV-K—these modified cells successfully killed breast cancer and melanoma cells in lab dishes and stopped tumor spread in animal studies.
For ALS, a separate team found that an antibody blocking HERV-K from connecting with its receptor on nerve cells reduced the neurotoxicity seen in patient samples.
HERV-K might actually be easier to target than HIV. The protein’s surface is less heavily coated with sugars that typically help viruses hide from the immune system. More of its vulnerable spots are exposed, particularly at the top where it likely binds to cells. That exposure could make it easier for therapeutic antibodies to latch on.
Why Dormant Viral DNA Wakes Up
HERV-K represents the most recently acquired viral DNA in the human genome. The genome contains roughly 100 full-length HERV-K sequences, though most picked up mutations over time that broke them. At least 10 locations still have intact genes capable of producing the envelope protein.
Healthy cells keep these viral remnants switched off through chemical modifications to DNA. In cancer and autoimmune disease, something disrupts that silencing mechanism. Cancer cells may reactivate HERV-K, and the protein is associated with increased cell growth and survival. In autoimmune conditions, HERV-K proteins can trigger inflammation, and when patients’ antibodies attack these proteins, the resulting immune complexes may contribute to disease progression.
The acidic environment inside tumors might favor the activated form of the protein, potentially explaining why it behaves differently in cancer versus autoimmune disease. Different versions of HERV-K in the genome produce proteins that stay trapped inside cells or reach the cell surface, which could account for varying disease patterns.
The study provides both a structural map of HERV-K and practical tools for detecting and targeting this protein. As cancer immunotherapy increasingly hunts for markers unique to tumor cells, these ancient viral proteins represent a largely unexplored target appearing across many cancer types. A virus that infected human ancestors millions of years ago has found new ways to cause problems today.
Disclaimer: The therapeutic approaches discussed involve laboratory and animal studies. These treatments have not yet been tested in human clinical trials.
Paper Notes
Limitations
The structures presented in this study used stabilized versions of the HERV-K envelope protein with engineered mutations, particularly the V437C and V498C cysteine pair that forms an artificial disulfide bond between subunits. While these modifications enabled structure determination, they may not perfectly represent the native protein’s behavior or dynamics. The postfusion structure excluded the hydrophobic fusion peptide (residues 466-497) and membrane-proximal region (residues 604-632) due to expression challenges. Some flexible regions, particularly the tether segment (residues 547-568) connecting HR1 and HR2 in the postfusion conformation, could not be fully modeled due to disorder. The antibodies were generated in mice and their epitopes on HERV-K may differ from human antibody responses in patients. The study focused on the consensus type 2 Phoenix envelope sequence, which may not capture variation among the multiple HERV-K loci present in the human genome.
Funding and Disclosures
This research received support from a Curebound discovery grant (13502-01-000-408) to E.O.S., funding from La Jolla Institute for Immunology & Kyowa Kirin Inc. (KKNA Kyowa Kirin North America) to E.O.S., and a Kyowa Kirin North America accelerator grant (18030-01-000-408) to J.S. The La Jolla Institute for Immunology has pending provisional patent coverage on the monoclonal antibodies and stabilized proteins described in this work. Patent applications entitled “HERV-K Envelope Protein Binders and Compositions and Methods of Use Thereof” (US Provisional Patent No. 63/638,067) and “Stabilized Pre-Fusion HERV-K Envelope Ectodomain Trimer” (US Provisional Patent No. 63/783,708) list inventors E.O.S., K.M.H., S.H., E.M.W., and J.S.
Publication Details
Authors: Jeremy Shek, Chen Sun, Elise M. Wilson, Fatemeh Moadab, Kathryn M. Hastie, Roshan R. Rajamanickam, Patrick J. Penalosa, Stephanie S. Harkins, Diptiben Parekh, Chitra Hariharan, Dawid S. Zyla, Cassandra Yu, Kelly C. L. Shaffer, Victoria I. Lewis, Ruben Diaz Avalos, Tomas Mustelin, Erica Ollmann Saphire
Affiliations: Center for Vaccine Innovation, La Jolla Institute for Immunology, La Jolla, CA; Department of Medicine, University of California San Diego, La Jolla, CA; Division of Rheumatology, University of Washington, Seattle, WA
Journal: Science Advances | Title: “Human endogenous retrovirus K (HERV-K) envelope structures in pre- and postfusion by cryo-EM” | DOI: 10.1126/sciadv.ady8168 | Publication Date: August 27, 2025 | Corresponding Author: Erica Ollmann Saphire ([email protected]) | Data Availability: Cryo-EM maps deposited in EM Database (EMD-70098, EMD-48351, EMD-48374); atomic models in Protein Data Bank (9O4F, 9MLA, 9MLK)







