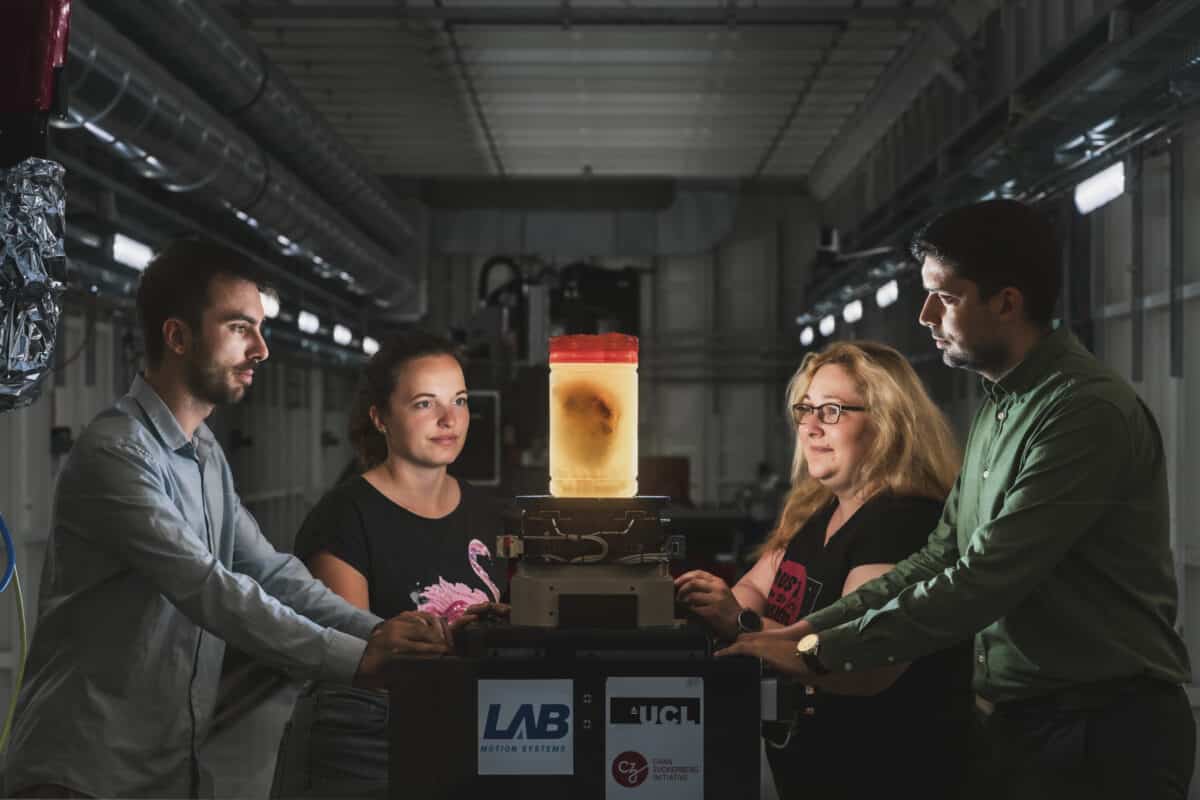

Joseph Brunet, UCL Senior Research Fellow and lead author, with Dheresa Urban, UCL Research Fellow, Joanna Purzycka, ESRF Engineer and Beamline Scientist, Hector Dejea I Velardo on ESRF Beamline BM18 preparing to scan a human heart. (Credit: ESRF/Stef Candé)
In A Nutshell
- Researchers have launched a free, publicly accessible 3D atlas of real human organs that lets anyone zoom from whole-organ views down to near-cellular detail in a web browser.
- The scans are made using a synchrotron, a massive particle accelerator that produces X-rays far more powerful than a hospital machine, and the imaging process leaves organs completely intact.
- The atlas currently holds 298 image sets from 24 donors across 10 organ types, including organs affected by COVID-19, cancer, hypertension, and rare conditions like Dandy-Walker syndrome.
- Early applications include training medical AI systems and giving anatomy students interactive 3D organ models built from real human tissue rather than idealized diagrams.
Pull up a web browser, type in an address, and within seconds a person can zoom from a satellite view of a continent down to the front door of a specific house. Now scientists have built something with the same basic logic, only the terrain is the human body. A new online atlas works something like a “Google Maps” for real human organs, letting anyone navigate in three dimensions, zooming from a whole heart down to individual muscle fibers.
Developed by researchers at University College London and the European Synchrotron Radiation Facility in Grenoble, France, the Human Organ Atlas is free and open to everyone, from surgeons to medical students to software engineers training artificial intelligence systems. No subscription, no institutional login, no specialized hardware required. Access is a web browser and a few clicks. Published in Science Advances, the project makes hundreds of organ scans available to the world at a level of detail that until recently existed only inside a handful of research laboratories.
A typical hospital CT scan can image an entire organ but with far lower detail than the new atlas. A microscope can reveal structures at near-cellular scale but only in a thin, flat slice of tissue that has been cut from the organ and destroyed in the process. Built on a technology called hierarchical phase-contrast tomography, or HiP-CT, the Human Organ Atlas covers both ends of that range and the scales in between, in three dimensions, all within a single intact organ.
The Technology Powering the Human Organ Atlas
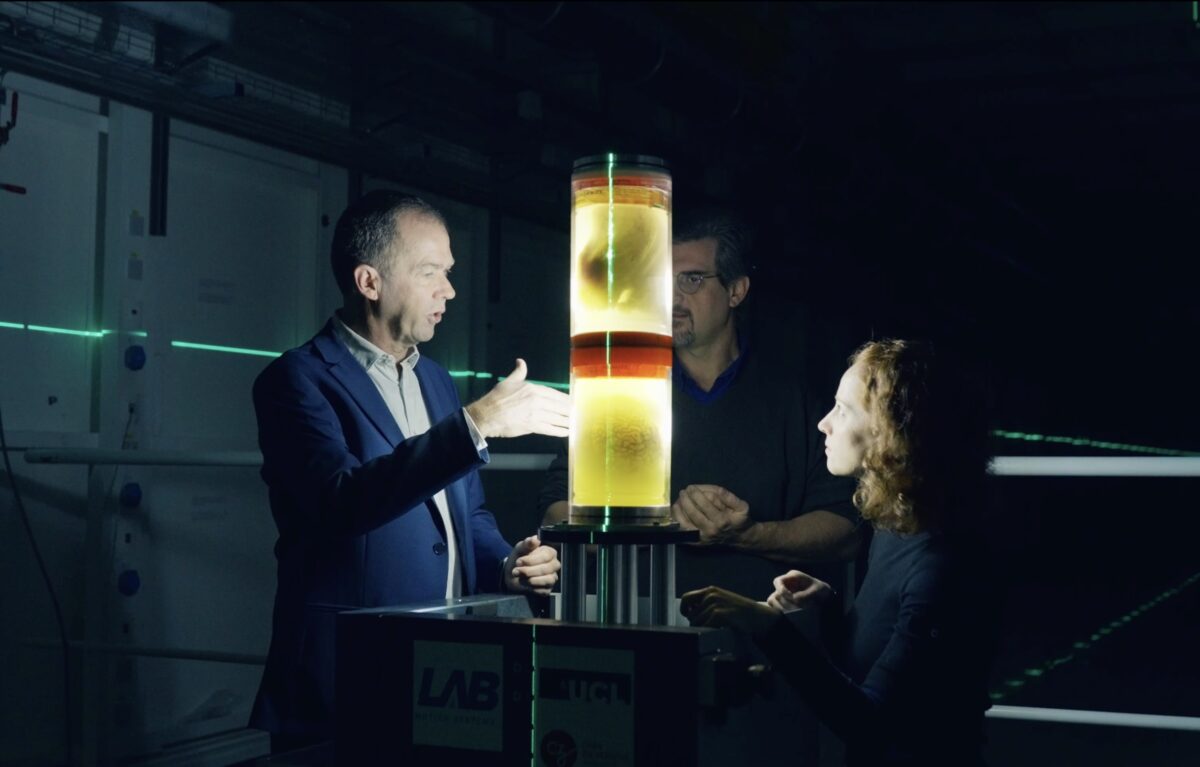
Getting there requires a machine most people will never encounter. A synchrotron is a particle accelerator the size of several city blocks. It generates X-ray beams far brighter and more tightly controlled than anything a hospital machine can produce, and the European Synchrotron Radiation Facility in Grenoble houses one of the most powerful such sources in the world.
Donated organs, sourced from European biobanks after death and with full donor consent, are preserved and mounted in the beam. A full-organ overview scan captures the entire sample at a resolution of roughly 20 micrometers per voxel (a voxel is the three-dimensional equivalent of a pixel). From there, the system zooms into selected regions at resolutions as fine as one micrometer per voxel, with the current record at 0.65 micrometers. All scans at every resolution are precisely aligned to one another, so moving between scales works exactly the way zooming works on a map. A full scanning session runs between two and eight hours depending on organ size.
The imaging itself does not require slicing the organ. Traditional histology, the standard method for seeing tissue at cellular-level detail, requires slicing an organ into sections thinner than a human hair. It is destructive and produces flat, two-dimensional images. HiP-CT leaves the organ physically intact from start to finish.

Inside the Human Organ Atlas: 10 Organs, 24 Donors
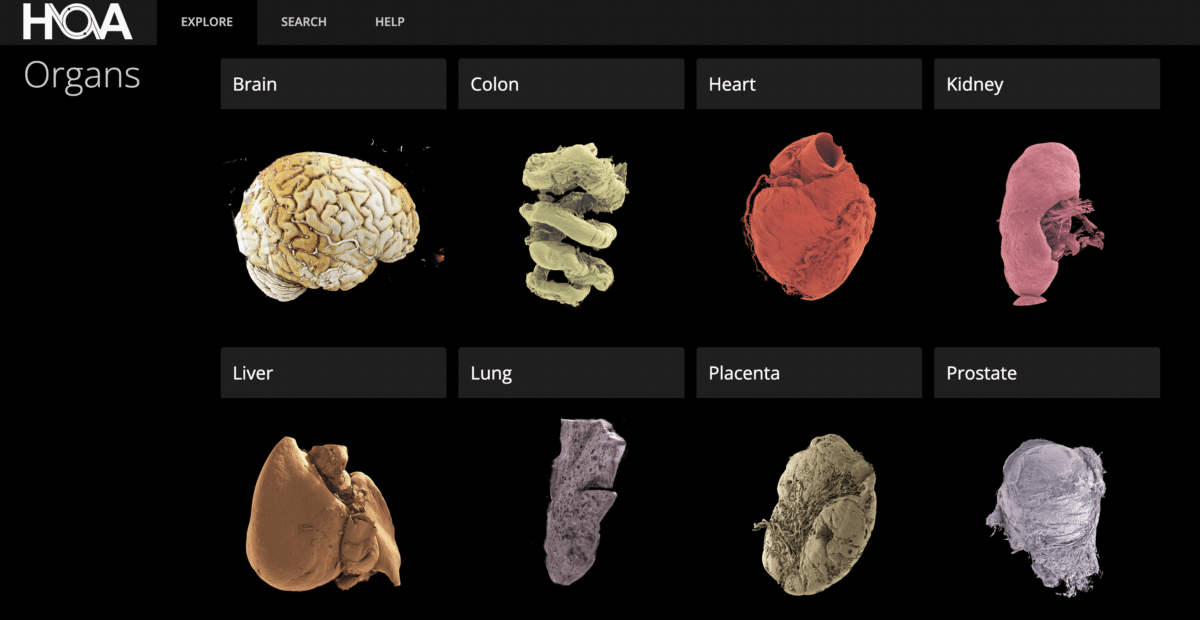
At the time of publication, the atlas holds 298 three-dimensional image sets from 24 individual donors, covering 10 organ types: brain, colon, heart, kidney, liver, lung, prostate, spleen, testis, and uterus. Donors range in age from 30 to 94, with an average age of 73. The collection currently skews older and male, reflecting who tends to donate bodies to European biobank programs. More than a dozen biobanks now contribute to the project, and future releases are expected to be more demographically varied.
Dataset sizes are enormous. Many exceed 200 gigabytes; some top one terabyte. To keep those files accessible to researchers without institutional supercomputers, the portal offers compressed, lower-resolution versions alongside the full data. A browser-based tool called neuroglancer lets anyone preview any scan interactively without downloading anything at all.
Beyond aging organs, the atlas holds a wide range of conditions. Many donors died during or after the COVID-19 pandemic, and the collection reflects that, with organs bearing evidence of hypertension, cancer, and COVID-19 damage. Rarer pathologies are represented too, among them a case of Dandy-Walker syndrome, a congenital brain malformation affecting fewer than one in 30,000 people. Most researchers would never have direct access to such a case through any other means.
How the Human Organ Atlas Is Already Changing Research and Education
Two early applications show what this kind of open access makes possible.
For artificial intelligence researchers, the atlas addresses a genuine shortage. Training AI systems to identify biological structures in three-dimensional medical images requires large volumes of high-quality data, and those datasets have been hard to come by. As a proof of concept, the research team used a process called super-resolution segmentation, which maps fine structures identified at high resolution onto lower-resolution whole-organ scans, to chart the tiny filtering units of the kidney known as glomeruli. Each glomerulus is between 100 and 300 micrometers across. Identifying and mapping all of them across an entire intact kidney, at scales not previously achievable, took a pipeline built on atlas data.
For medical educators, the atlas offers something anatomy textbooks and cadaver labs cannot: real, disease-bearing human organs in fully navigable three-dimensional detail. Using a technique called 3D Gaussian splatting, which compresses large three-dimensional datasets into lightweight, interactive displays, organ data can be rendered on standard consumer hardware. Students can rotate a heart, zoom into a valve, and descend further still into the microscopic fiber arrangement of the tissue beneath it, all from a single continuous dataset drawn from an actual human organ.
Neither application is theoretical. Both are running on data in the atlas today.
A person exploring the Human Organ Atlas for the first time might start with a whole kidney, follow the branching arteries across its surface, zoom into a cluster of vessels, and descend into the tissue where the filtering structures sit, all within a browser window, all from a real organ that once belonged to a real person. That is not a model or a simulation. It is the body, mapped.
Paper Notes
Limitations
Because all organs were imaged after death rather than in living subjects, several artifacts are present in the data. Organ shrinkage from the preservation process is visible in most datasets, and large internal chambers, particularly the left atrium of the heart, appear collapsed. Blood vessels are frequently compressed, posing challenges for anyone mapping vascular structures. High-resolution scanning introduces a theoretical risk of radiation damage to samples, though the team states the HiP-CT acquisition protocol was built to prevent this. Donor demographics at publication skew older and male, limiting how broadly representative the current dataset is. Medical history for individual donors, collected through external biobanks, cannot be assumed complete.
Funding and Disclosures
Funding was provided by the Chan Zuckerberg Initiative, the NIH BRAIN Initiative CONNECTS program through the National Institute of Neurological Disorders and Stroke and the National Institute of Mental Health, the Wellcome Trust, the NIHR UCLH Biomedical Research Centre, Siemens Healthineers AG, the special research fund of the University of Antwerp, the Collen-Francqui Foundation, the Royal Academy of Engineering, and a CIFAR MacMillan Fellowship. Klaus Engel receives wages from Siemens Healthineers AG and holds stock in Siemens entities. Joseph Jacob declares consultancy and lecture fees from Boehringer Ingelheim, F. Hoffmann-La Roche, GlaxoSmithKline, and Takeda, as well as grant funding from GlaxoSmithKline, Microsoft Research, and the Chan Zuckerberg Initiative, among others. All other authors declare no competing interests.
Publication Details
Title: The Human Organ Atlas Authors: Claire L. Walsh, Joseph Brunet, David Stansby, Guillaume Gaisné, Yang Zhou, Maximilian Ackermann, Alexandre Bellier, Camille Berruyer, Axel Bocciarelli, Marjolaine Bodin, Bernadette S. de Bakker, Hector Dejea, Alejandro De Maria Antolinos, Klaus Engel, Andy Götz, Joseph Jacob, Daniel Jonigk, Joanna Purzycka, Theresa Urban, Stijn E. Verleden, Ruikang Xue, Paul Tafforeau, Peter D. Lee Journal: Science Advances, Vol. 12, Issue 11, eadz2240 Published: March 11, 2026 DOI: 10.1126/sciadv.adz2240







