
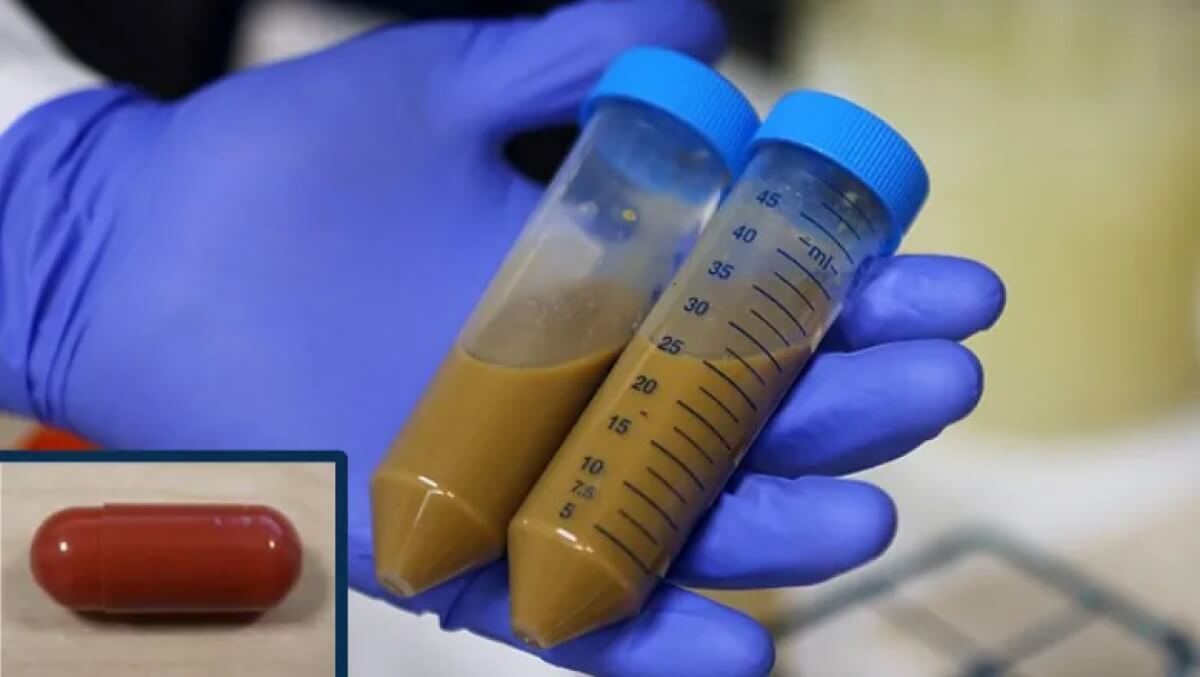
Fecal transplant container and pills. (© TopMicrobialStock - stock.adobe.com)
COPENHAGEN, Denmark — Just as modern vaccines evolved from cowpox blisters and penicillin from moldy bread, another life-saving treatment is undergoing a remarkable transformation. Scientists have developed a novel method to cultivate beneficial viruses from human feces, potentially revolutionizing how we treat gastrointestinal disorders and other health conditions.
You’ve probably heard of fecal transplants – an increasingly common medical procedure where doctors transfer stool from a healthy donor to a sick patient. Despite sounding unsettling, the treatment has shown remarkable success in treating certain gut infections, particularly those caused by a nasty bacteria called Clostridioides difficile (better known as C. diff). But there’s a catch: transferring someone else’s stool carries risks, including the possible transmission of harmful viruses or bacteria that might have slipped through screening.
The human gut hosts approximately 100 trillion microorganisms, including both beneficial and potentially harmful species. This complex ecosystem profoundly influences our overall health, affecting everything from our immune response to our mental well-being. When this system becomes unbalanced, various health issues can emerge. Traditional fecal transplants help restore this balance by introducing a complete microbial community from a healthy donor, but the new method offers a more refined approach.
Enter the heroes of our story: bacteriophages, or “phages” for short. These are specialized viruses that only infect bacteria, not human cells. Think of them as nature’s bacterial assassins, evolved over millions of years to keep bacterial populations in check. Scientists at Tallinn University of Technology and the University of Copenhagen have developed a clever way to harness these bacterial predators while minimizing the risks associated with traditional fecal transplants.
This advancement could transform a treatment that currently relies on direct fecal transfers into a standardized, pharmaceutical-grade therapy. The new technique, published in the journal iScience, addresses these limitations by using fermentation technology – the same basic process used to make foods like kombucha and sourdough bread – to cultivate specific beneficial components from fecal matter.
The researchers employed a sophisticated device called a chemostat – essentially a controlled fermentation tank – to create ideal growing conditions for beneficial gut microorganisms. The system continuously adds fresh nutrient solutions while removing waste products, allowing researchers to maintain precise control over the microbial environment. This process gradually washes away potentially harmful viruses while maintaining and even enriching the population of beneficial bacteriophages.
After running the system for about five days, the relative abundance of viruses that could potentially infect human cells dropped to less than 0.01% of the total viral population – a hundred-fold reduction from the starting material. Meanwhile, the beneficial phages maintained their diversity and stability, creating reproducible communities that could potentially be used as therapeutic tools.
This cultivation method represents a significant step toward standardizing gut microbiome treatments. Current fecal transplants, while effective, are something of a “black brown box” – medical professionals can’t precisely specify what’s being transferred from donor to recipient. The new technique could allow for consistent, reproducible treatments where doctors know exactly what they’re administering to patients.
Looking ahead, the researchers envision developing precisely formulated pills that could be prescribed by doctors or purchased at pharmacies. The treatment’s potential extends beyond just gastrointestinal disorders – research suggests that gut health plays a crucial role in conditions ranging from asthma to Type 1 diabetes and even mental health.
The commercial potential of this advancement is significant. Current fecal transplant procedures require extensive (and expensive) screening of donors, limiting treatment availability. A standardized, cultivated alternative could dramatically increase accessibility while potentially expanding the range of treatable conditions.
“Ultimately, our hope is to have a product that is free of bacteria and potentially harmful viruses, leaving only a purified dose of good viruses, called bacteriophages, that can potentially fight various gastrointestinal disorders and, in the long term, be used to treat a wide range of health issues,” says Torben Sølbeck Rasmussen, an assistant professor from the University of Copenhagen’s Department of Food Science, in a statement.
The evolution of this treatment mirrors the development of other revolutionary medical advances. Just as we no longer need to harvest cowpox blisters for vaccines or grow penicillin from bread mold, the future of gut microbiome therapy may not require direct fecal transfers. Instead, it may come in the form of carefully cultivated beneficial viruses, offering a cleaner, safer, and more standardized approach to treating a wide range of health conditions.
Paper Summary
Methodology
The researchers used a specialized device called a chemostat to continuously culture gut microbes and their associated viruses. They started with two types of samples: pooled cecal (gut) contents from mice and pooled fecal samples from healthy human donors. The mouse samples came from 18 different mice of the same strain but from different vendors to ensure diversity. The human samples came from seven healthy donors aged 19-37 years.
These samples were placed in the chemostat under carefully controlled conditions that mimicked the gut environment, including specific temperature, pH levels, and nutrient mixtures. The system ran continuously, with fresh medium flowing in and waste flowing out at two different speeds (called dilution rates) to simulate different gut transit times. They ran each experiment for five “residence times” – the time it takes to completely replace the culture volume – and collected samples throughout the process to analyze the bacterial and viral communities.
Key Results
The main finding was that their chemostat system successfully reduced the relative abundance of eukaryotic viruses (those that can infect human cells) from about 0.2% to below 0.01% in most cases. Meanwhile, the beneficial bacteriophages maintained their diversity and showed reproducible patterns between different experiments.
The system produced about 2 billion virus-like particles per milliliter, with eukaryotic viruses making up less than 100,000 of these – potentially below the threshold needed to cause infection. The bacterial communities also showed stable and reproducible patterns, with different species thriving at different flow rates.
Study Limitations
First, the researchers’ laboratory setup cannot fully replicate the complex conditions of the real gut environment. Second, while they could track the reduction in eukaryotic viruses, they need more specific methods to monitor particular virus types. Third, some of their growth medium components came from animal and plant sources, which might have introduced some viral particles. Finally, there’s also a possibility that some viruses might have been misclassified due to similarities between viral genes.
Discussion & Takeaways
The study demonstrates a promising new approach to creating therapeutic viral communities with minimal risks. The method could potentially replace or complement traditional fecal transplants, offering a more controlled and reproducible treatment option.
The researchers have already tested their approach in treating C. difficile infections in mice, showing comparable success rates to traditional treatments. The system’s ability to maintain diverse and stable viral communities while washing out potentially harmful viruses represents a significant step forward in developing microbiome-based therapies.
Funding & Disclosures
The research was funded by several organizations, including the Estonian Ministry of Science and Education, Tallinn University of Technology, and the Lundbeck Foundation. The Novo Nordisk Foundation also provided support under the project name “PrePhage.” The authors declared no competing interests in their research.







